We live in a world that is susceptible to emergence of new infectious diseases due to climate change and inter-connectedness among countries like never before. COVID-19 has been the most recent one. And our memories are fresh with many other epidemics in the last two decades. Given the connectedness of the modern world, infectious diseases spread throughout the world rather easily. While our history is replete with numerous epidemics and pandemics in which millions of lives have been lost, they have also taught us the importance of better sanitation, improving infrastructure for health care, holding large scale vaccination drives to prevent future outbreaks. They have also taught the importance of detecting the infection spreads as early as possible. The sooner we detect the emergence or spread of a particular infection, the better prepared we can be to fight them and prevent losses of lives and other resources. This is where the importance of disease surveillance comes in.
Surveillance of a disease is keeping a track of its spread. It is primarily done manually where every hospital reports the number of cases they received of a particular disease. Large scale organisations such as World Health Organisation (WHO), Centre for Disease Control and Prevention (CDC) have extensive databases and monitor several diseases through established large-scale networks. Disease surveillance by these methods are most helpful in monitoring and tracking an ongoing epidemic. But one can also track the causative micro-organisms of the diseases. Their surveillance in metropolitan cities, the hotspots of infection spread, using wastewater is one of the clever ways by which cities can always be prepared to act against an emerging epidemic.
This form of surveillance is not new; it was first carried out in the 1940s in the United States to track and contain the poliovirus outbreak. In the last two years, some of the other countries have used this to track and monitor COVID-19. The CSIR-Centre for Cellular and Molecular Biology (CCMB) at Hyderabad, India is one such centre where they have successfully established a wastewater surveillance program for monitoring of COVID-19 in India. Initially started in Hyderabad, this program has now expanded its reach to several cities in Maharashtra, as well as Kolkata, Delhi and Chennai. To gain more insight about how this program operates, I (KB) decided to meet Dr Archana Bharadwaj Siva (ABS). She is a senior scientist who leads the wastewater surveillance program at CCMB.
Here are some excerpts from our discussion.
—————————————————————————————————————
KB: What is wastewater surveillance, and how does it help in infectious disease management?
ABS: One of the applications of wastewater surveillance is basically tracking pathogens that cause infectious diseases (as is being done at CCMB presently) and its spread across a region by monitoring the sewage of the region. This method is highly effective in detecting pathogens that are shed through faeces, such as SARS-COV-2 (causative pathogen for COVID-19). Sewage or wastewater is collected from different wastewater treatment plants throughout the locality and presence of the pathogen is detected through several approaches such as PCR, rt-PCR (Reverse Transcriptase PCR), qRT-PCR (quantitative Real Time PCR) or antibody tests. All of these help in detecting the genetic material or the proteins of the pathogens.) This method helps us in tracking a disease for days, even months after its spread and also gives us an idea about how much the disease has spread throughout the population.
KB: Can you please elaborate the steps followed between collecting water samples and predicting the spread of SARS-CoV-2?
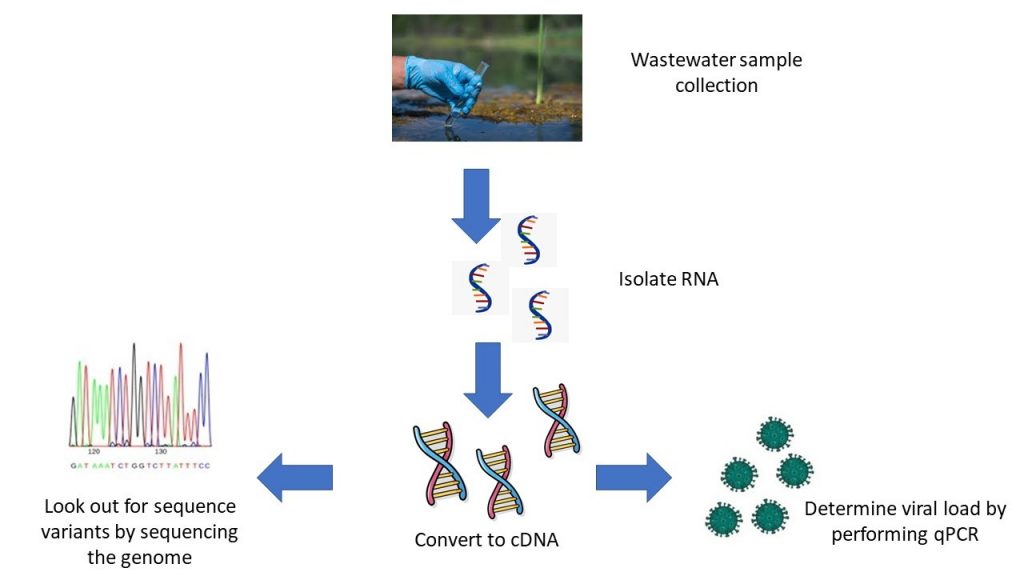
ABS: We first collect sewage samples from different wastewater treatment plants, or open flowing nallahs/drains and isolate RNA from these samples (since RNA is the genetic material of SARS-CoV-2). But the isolate contains RNA of all kinds of living things in the sample. For tracking SARS-CoV-2, we look for RNA of the coronavirus specifically. We first detect the presence and quantitate the viral load through real time RT-PCR. Then we attempt identifying the variant by sequencing the genome in a way that we selectively choose for the coronavirus genome. Not only that, it also helps in tracking the virus’ variants, old and new. It also hints at which variant is prevalent in a given population. We also quantitate the viral load of the site by performing qRT-PCR using the isolated RNA.

KB: Can you please give an example of how this method of disease surveillance has helped any of the Indian cities in making informed decisions about infection control?
ABS: We are yet to see many examples from India. However, from our analysis of one of the regions, we had observed that there was a spike in SARS-CoV-2 load in wastewater approximately two weeks before the COVID-19 cases began to surge in the month of June 2022. There have been cases abroad where wastewater surveillance has given an almost 4 weeks forewarning about the spread of infection, thus, helping cities in preparing for an emerging outbreak.
KB: Now that this system has been well-established for surveillance of COVID-19, could this system also be used to monitor other diseases in India?
ABS: Any pathogen that is shed through faeces and is stable for a fairly long period of time can be monitored using wastewater surveillance. The National Centre for Disease Control (NCDC) has been routinely monitoring poliovirus through wastewater surveillance under its Integrated Disease Surveillance Program (IDSP).
Talking of the scope of this technology, we are definitely thinking of tracking antimicrobial resistance (AMR) profiles using this method. There is increasing evidence of rising drug resistance (including towards antibiotics) in several strains responsible for many infectious diseases, such as for TB. These strains are difficult to control using the available antimicrobial drugs, which makes the disease harder to cure. It is possible to identify strains of antimicrobial resistant bacteria, viruses or other microbes through wastewater surveillance provided these microbes are shed through bodily waste.
We are developing a system for understanding AMR profiles in localities by first performing untargeted genome sequencing on their wastewater. This will help us know if there are antimicrobial resistant strains/genes present in the locality without needing to identify the exact strain at first. Unlike RNA isolation in COVID-19 surveillance, here we isolate genomic DNA from the sewage samples we collect from the sewage treatment plants or open drains/nallahs since DNA is the genetic material of most microbes. The genomic DNA is sequenced, and their sequences are then matched with a database of antimicrobial resistant strains to identify any of the resistant strains present in the sample.
If we detect an antimicrobial resistance profile, then we will proceed to targeted genome sequencing. That will help us look for strains of a particular bacteria, for example, different strains of Mycobacterium tuberculosis.
Such information would be useful to track the prevalence of a particular antimicrobial-resistant strain in that given population, and could help healthcare agencies in devising different management strategies.
—————————————————————————————————————
Innovative ways of infection control to limit its spread throughout the population are always sought after, and surveillance of diseases using wastewater has been a successful approach. This method has reached a policymaking stage in India, where every state can consider to incorporate this approach for tracking and surveillance of different diseases in the community. This way states would be better prepared to fight off many diseases and epidemics as soon as they emerge.


